krogan lab
cardiovascular disease
Systems Analysis for Understanding Complex Biological Phenomena
Our laboratory employs global approaches to gain insight into a wide variety of biological systems. We are primarily focused on generating and analyzing genetic and protein-protein interaction maps. Using these datasets, we aim to derive mechanistic insights into how specific pathways, complexes and proteins function in various processes. This “systems to mechanism” approach is being applied to a variety of organisms, including bacteria, yeast and mammalian cells, as well as pathogenic agents.
To characterize physical interactions, we use systematic affinity tag-purification/mass spectrometry (AP-MS) to identify the protein complexes in a given organism or process. Then, use of genetic interaction data allows us to place the protein complexes into functional pathways.
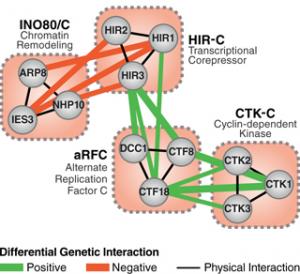
We developed a quantitative genetic approach to determine their dependency of functionally related sets of genes, termed E-MAP, and applied this strategy to Saccharomyces cerevisiae, Schizosaccharomyces pombe and mammalian cells. This combined physical and genetic interrogation has been used to study a wide variety of biological processes, including DNA repair, chromatin function, protein trafficking, RNA processing and signaling, gaining insights into pathway organization all the way to the function of individual residues on specific proteins. We also study how these networks change over time, which can be fast (e.g., response to a stimulus) or slow (e.g., speciation). For example, we have been generating genetic interaction maps under different conditions (termed differential E-MAP, or dE-MAP), such as in the presence of DNA-damaging agents or specific stresses, which has allowed us to identify individual pathways that only function under given stresses. Furthermore, our group also studies the conservation of these networks across different species, which provides insight into evolution and the molecular mechanisms involved in speciation. We are now using the E-MAP and AP-MS approaches to determine the critical networks that function during cardiac differentiation and are disrupted in human cardiac disease.
In nature, organisms often interact with other organisms and we have also been using systems approaches to understand these inter-species relationships. For example, using AP-MS and E-MAP, we have been studying the physical and genetic interface between pathogenic organisms, including tuberculosis, dengue, human papillomavirus and HIV and their hosts. This work is leading to a more comprehensive understanding of how pathways are hijacked and re-wired during the course of infection. Finally, we are functionally interrogating different sets of genes that have been linked to various disease states, including heart disease and cancer. Generating a more holistic view of these networks, and how they are perturbed in the presence of specific mutations, will provide insight into the molecular mechanisms that are responsible for these different diseases.


